The xarray.Dataset structure for AITA#
xarrayaita is a dataset accessor for xarray. In this section we explain what is the minimal construction you have to do and what functions you can apply through this xarrayaita
The minimal structure of xarrayaita#
import xarrayaita.loadData_aita as lda #here are some functions to build the xarrayaita structure
import xarrayaita.aita as xa
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import xarray as xr
Load data to show an exemple.
# path to data and microstructure
path_data='orientation_test.dat'
path_micro='micro_test.bmp'
In this case we use a constructor function that takes as input :
a
.datfile which is an output from G50 fabric analyser. If you have another analyser it should be easy to implement another constructor function.a
.bmpwhich is the microstructure image (black and white, obtained from a segmentation for instance) that must have been extracted elsewhere to get the grain boundaries from the orientation image (grain boundaries are the white pixels). This input can be left empty
help(lda.aita5col)
Help on function aita5col in module xarrayaita.loadData_aita:
aita5col(adr_data, micro_adress=0)
Function to load the data from G50 analyser that have 5 columns
:param adr_data: path to the data
:type adr_data: str
:param micro_adress: path to microstructure file (black and white image where grains boundaries are white) `.bmp`
:type micro_adress: str
data=lda.aita5col(path_data,path_micro)
xarray.Dataset#
The data structure is a regular xarray.Dataset. Therefore it can be used as one. (Pas clair!)
Some specific functions can be applied to it by:
data.xa.FUNCTION
You can visualise what is stored in your xarray.Dataset using:
data
<xarray.Dataset>
Dimensions: (uvecs: 2, x: 1000, y: 2500)
Coordinates:
* x (x) float64 0.0 0.02 0.04 0.06 0.08 ... 19.92 19.94 19.96 19.98
* y (y) float64 49.98 49.96 49.94 49.92 49.9 ... 0.06 0.04 0.02 0.0
Dimensions without coordinates: uvecs
Data variables:
orientation (y, x, uvecs) float64 2.395 0.6451 5.377 ... 0.6098 0.6473
quality (y, x) int64 0 90 92 93 92 92 94 94 ... 96 96 96 96 96 97 97 96
micro (y, x) float64 0.0 0.0 0.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0 0.0
grainId (y, x) int64 1 1 1 1 1 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1 1 1 1
Attributes:
date: Thursday, 19 Nov 2015, 11:24 am
unit: millimeters
step_size: 0.02
path_dat: orientation_test.dat- uvecs: 2
- x: 1000
- y: 2500
- x(x)float640.0 0.02 0.04 ... 19.94 19.96 19.98
array([ 0. , 0.02, 0.04, ..., 19.94, 19.96, 19.98])
- y(y)float6449.98 49.96 49.94 ... 0.04 0.02 0.0
array([4.998e+01, 4.996e+01, 4.994e+01, ..., 4.000e-02, 2.000e-02, 0.000e+00])
- orientation(y, x, uvecs)float642.395 0.6451 ... 0.6098 0.6473
array([[[2.39476627, 0.64507369], [5.37718489, 1.04999008], [5.38905313, 1.05627326], ..., [2.05826679, 0.49654617], [5.65731024, 0.94160513], [5.68523551, 1.03218772]], [[5.35955707, 1.15837502], [5.35885894, 1.12975162], [5.3613024 , 1.11701072], ..., [2.07659274, 0.52063172], [5.64753639, 0.94073247], [5.67912685, 1.04143796]], [[5.36304773, 1.23045712], [5.36165146, 1.23132979], [2.38656322, 0.57246799], ..., ... ..., [0.62378067, 0.625526 ], [0.61784656, 0.61994095], [0.61645029, 0.61400683]], [[0.68085294, 1.12399204], [0.68050388, 1.12730816], [0.67840948, 1.13900187], ..., [0.64123397, 0.62954026], [0.62430427, 0.63529985], [0.61575216, 0.62203535]], [[0.67701322, 1.13236962], [0.67753682, 1.13690747], [0.67840948, 1.13149695], ..., [0.63652158, 0.61418136], [0.61645029, 0.63948864], [0.60981804, 0.64734262]]]) - quality(y, x)int640 90 92 93 92 92 ... 96 96 97 97 96
array([[ 0, 90, 92, ..., 0, 85, 90], [81, 82, 83, ..., 0, 84, 89], [81, 80, 3, ..., 0, 79, 88], ..., [90, 91, 91, ..., 95, 95, 95], [91, 92, 91, ..., 95, 95, 95], [92, 92, 91, ..., 97, 97, 96]]) - micro(y, x)float640.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
array([[0., 0., 0., ..., 0., 0., 0.], [0., 0., 0., ..., 0., 0., 0.], [0., 0., 0., ..., 0., 0., 0.], ..., [0., 0., 0., ..., 0., 0., 0.], [0., 0., 0., ..., 0., 0., 0.], [0., 0., 0., ..., 0., 0., 0.]]) - grainId(y, x)int641 1 1 1 1 1 1 1 ... 1 1 1 1 1 1 1 1
array([[1, 1, 1, ..., 1, 1, 1], [1, 1, 1, ..., 1, 1, 1], [1, 1, 1, ..., 1, 1, 1], ..., [1, 1, 1, ..., 1, 1, 1], [1, 1, 1, ..., 1, 1, 1], [1, 1, 1, ..., 1, 1, 1]])
- date :
- Thursday, 19 Nov 2015, 11:24 am
- unit :
- millimeters
- step_size :
- 0.02
- path_dat :
- orientation_test.dat
There are 4 xarray.DataArray variables :
orientation: stores the orientation of each pixel as azimuth and colatitude (see figure below). It is meant to be used with xarrayuvecsDataArray.quality: the quality factor given by the analyser (between \([0,100]\)).micro: the binary microstructure image given as an input.grainId: label for each grain of the microstructure. Each pixel of a grain as a givengrainIdlabel.
This is the minimal structure necessary to use the xarrayaita function.
Visualisation of the variables#
Every Data variable can be plotted using a regular xarray plot function
Orientation variable#
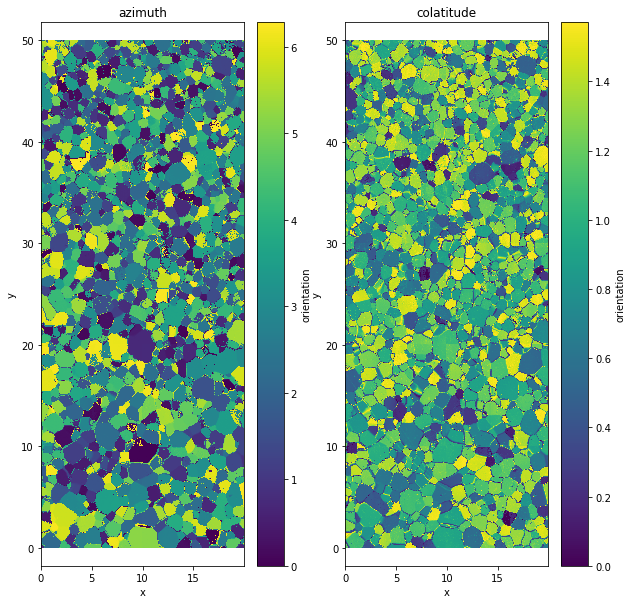
The data are stored as two angles, azimuth and colatitude
plt.figure(figsize=(10,10))
plt.subplot(1,2,1)
data.orientation[:,:,0].plot()
plt.axis('equal')
plt.title('azimuth')
plt.subplot(1,2,2)
data.orientation[:,:,1].plot()
plt.axis('equal')
plt.title('colatitude')
Text(0.5, 1.0, 'colatitude')
Quality variable#
plt.figure(figsize=(5,10))
data.quality.plot(cmap=cm.gray)
plt.axis('equal')
plt.title('quality')
Text(0.5, 1.0, 'quality')

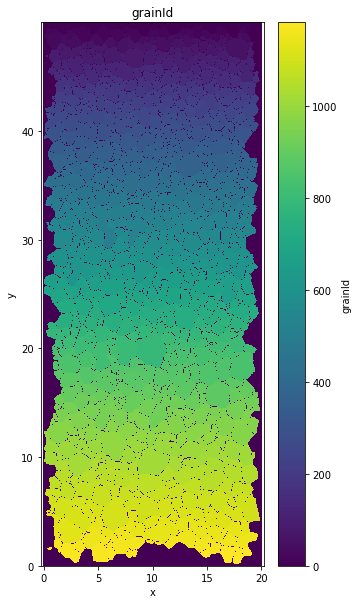
GrainID variable#
plt.figure(figsize=(5,10))
data.grainId.plot()
plt.axis('equal')
plt.title('grainId')
Text(0.5, 1.0, 'grainId')
If you prefer to look at Bunge Euler angles you can extract them using uvecs
N.B. : TO DO, say how
All the variables are visible in your xarray.Dataset
Functions within xarrayaita#
xa.filter#
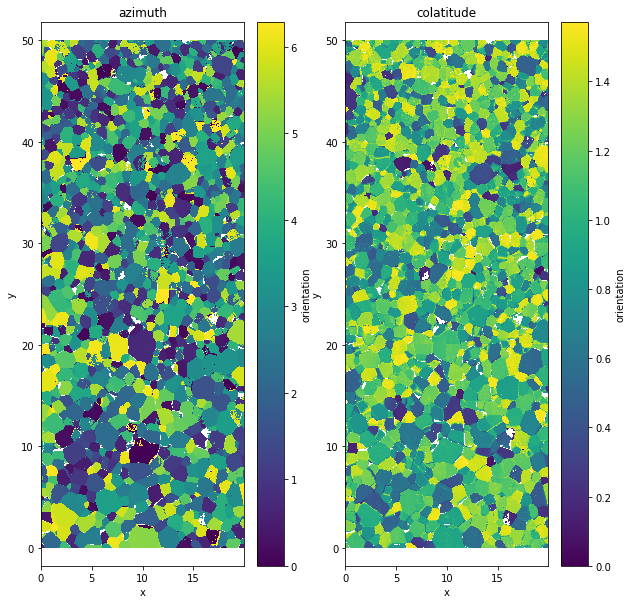
It only affects the orientation value. When the data.quality variable is below the given value val the orientation value is replaced by a nan.
help(xa.aita.filter)
Help on function filter in module xarrayaita.aita:
filter(self, val)
Put nan value in orientation file
data.aita.filter(75)
plt.figure(figsize=(10,10))
plt.subplot(1,2,1)
data.orientation[:,:,0].plot()
plt.axis('equal')
plt.title('azimuth')
plt.subplot(1,2,2)
data.orientation[:,:,1].plot()
plt.axis('equal')
plt.title('colatitude')
Text(0.5, 1.0, 'colatitude')
Symmetry transformation#
fliplr#
Flip the data with a horizontal mirror.
help(xa.aita.fliplr)
Help on function fliplr in module xarrayaita.aita:
fliplr(self)
flip left right the data and rotate the orientation
May be it is more a routation around the 0y axis of 180 degree
rot180#
help(xa.aita.rot180)
Help on function rot180 in module xarrayaita.aita:
rot180(self)
rotate 180 degre around Oz the data and rotate the orientation
rot90c#
help(xa.aita.rot90c)
Help on function rot90c in module xarrayaita.aita_geom:
rot90c(self)
Geometric transformation
Rotate 90 degre in clockwise direction
Return :
- xarray.Dataset : rotated dataset
